05: Pandas
Reality check: Data scientists spend 80% of their time cleaning data and 20% complaining about it. The remaining 20% is spent on actual analysis (yes, that’s 120% - data science is just that intense!)
 Shows the reality that data cleaning is most of the work - perfect intro to data cleaning lecture
Shows the reality that data cleaning is most of the work - perfect intro to data cleaning lecture
Data cleaning follows a systematic workflow: detect → handle → validate → transform. We’ll cover each technique individually, then bring it all together in a complete pipeline at the end.
Handling Missing Data
Section titled “Handling Missing Data”Missing data is a common problem in real-world datasets. Understanding how to identify, analyze, and handle missing data is crucial for reliable data analysis. Pandas provides powerful tools for working with missing values.
Fun fact: Missing data has its own Wikipedia page with 47 different types of missingness. The most common? “I forgot to fill this out” and “The system crashed again.”
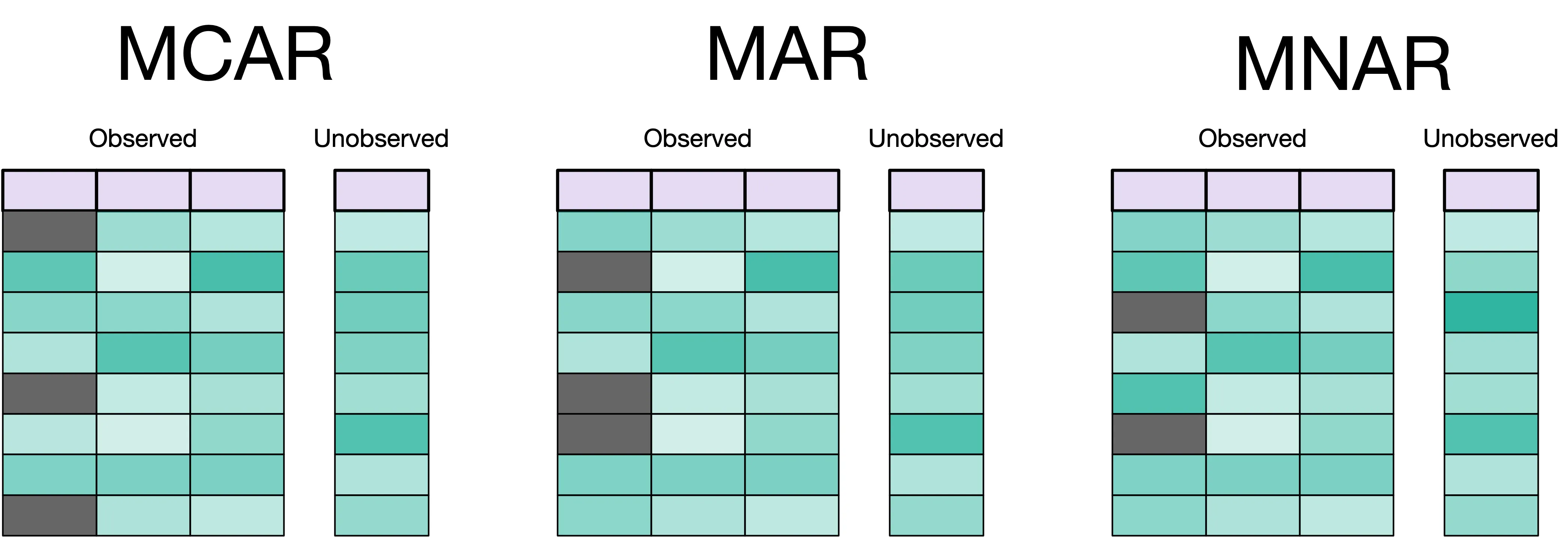
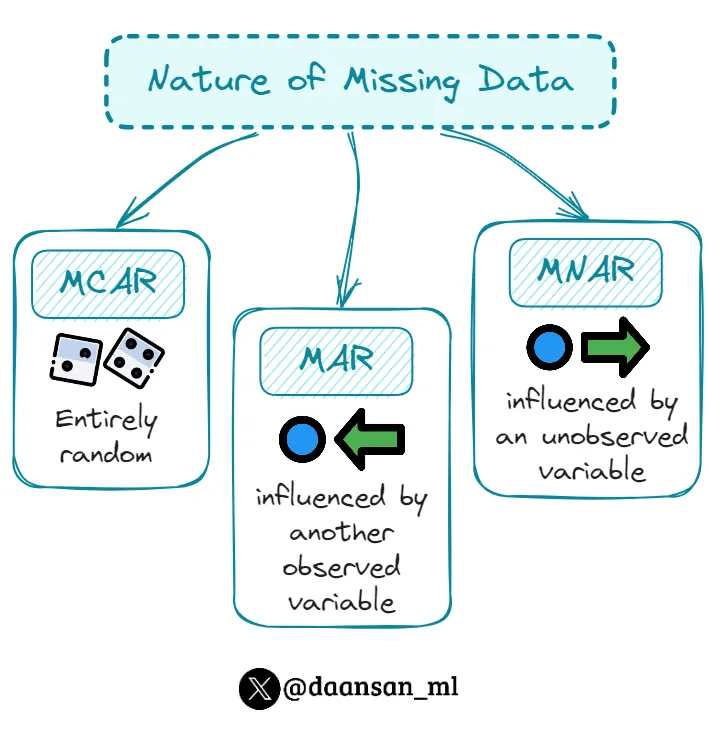 Common missing data patterns: MCAR (Missing Completely At Random), MAR (Missing At Random), MNAR (Missing Not At Random)
Common missing data patterns: MCAR (Missing Completely At Random), MAR (Missing At Random), MNAR (Missing Not At Random)
Missing Data Detection
Section titled “Missing Data Detection”Missing data detection identifies where data is missing and helps understand the pattern of missingness. This is the first step in any data cleaning process.
Pro tip: Missing data is like that one friend who’s always late to everything - you know they’re supposed to be there, but you can never quite predict when (or if) they’ll show up.
Reference:
df.isnull()- Boolean DataFrame: True for missing valuesdf.notnull()- Boolean DataFrame: True for non-missing valuesdf.isna()- Alias for isnull()df.notna()- Alias for notnull()df.isnull().sum()- Count missing values per columndf.isnull().any()- True if any missing values in columndf.isnull().all()- True if all values missing in column
Example:
# Check for missing valuesdf = pd.DataFrame({'A': [1, 2, None, 4], 'B': [5, None, 7, 8]})print(df.isnull().sum()) # A: 1, B: 1print(df.isnull().any()) # A: True, B: Trueprint(df.isnull().all()) # A: False, B: False
# Visualize missing dataimport matplotlib.pyplot as pltdf.isnull().sum().plot(kind='bar')plt.title('Missing Values by Column')plt.show()Missing Data Analysis
Section titled “Missing Data Analysis”Missing data analysis helps understand the pattern and mechanism of missingness. This information guides the choice of appropriate handling strategies.
Reference:
df.isnull().sum()- Count missing values per columndf.isnull().sum(axis=1)- Count missing values per rowdf.isnull().mean()- Proportion of missing values per columndf.dropna()- Remove rows with any missing valuesdf.dropna(axis=1)- Remove columns with any missing valuesdf.dropna(thresh=n)- Keep rows with at least n non-null values
Example:
# Analyze missing data patternsdf = pd.DataFrame({'A': [1, 2, None, 4], 'B': [5, None, 7, 8], 'C': [9, 10, 11, None]})print(df.isnull().sum()) # Missing values per columnprint(df.isnull().mean()) # Proportion missing per columnprint(df.isnull().sum(axis=1)) # Missing values per row
# Remove rows with missing valuesdf_clean = df.dropna()print(df_clean.shape) # (2, 3) - removed rows with missing valuesMissing Data Imputation
Section titled “Missing Data Imputation”Missing data imputation fills in missing values using various strategies. The choice of imputation method depends on the data type and the pattern of missingness.
Reference:
df.fillna(value)- Fill missing values with constantdf.ffill()/df.fillna(method='ffill')- Forward fill (use previous value; method= deprecated in pandas 3.0)df.bfill()/df.fillna(method='bfill')- Backward fill (use next value; method= deprecated in pandas 3.0)df.fillna(df.mean())- Fill with column meandf.fillna(df.median())- Fill with column mediandf.fillna(df.mode().iloc[0])- Fill with column modedf.interpolate()- Interpolate missing values
Example:
# Fill missing valuesdf = pd.DataFrame({'A': [1, 2, None, 4], 'B': [5, None, 7, 8]})
# Fill with constantdf_filled = df.fillna(0)print(df_filled) # Missing values replaced with 0
# Fill with meandf_mean = df.fillna(df.mean())print(df_mean) # Missing values replaced with column mean
# Forward fill - modern syntax (pandas 1.4+)df_ffill = df.ffill()print(df_ffill) # Missing values replaced with previous value
# Deprecated syntax (still works in pandas 2.x, removed in 3.0)df_ffill_old = df.fillna(method='ffill') # Avoid this in new codeOriginal Data: Forward Fill (ffill): Backward Fill (bfill): Index Value Index Value Index Value 0 10 0 10 ─┐ 0 10 1 [NaN] 1 10 ←┤ fills down 1 15 ←┐ 2 [NaN] 2 10 ←┘ from 10 2 15 ←┤ fills up 3 15 3 15 ─┐ 3 15 ─┘ from 15 4 [NaN] 4 15 ←┤ fills down 4 [NaN] can't fill 5 [NaN] 5 15 ←┘ from 15 5 [NaN] no later rowsLIVE DEMO! (Demo 1: Missing Data - detection, analysis, and imputation strategies)
Data Transformation Techniques
Section titled “Data Transformation Techniques”Removing Duplicates
Section titled “Removing Duplicates”Duplicate rows can skew your analysis and waste computational resources. Removing duplicates is a common first step in data cleaning.
Fun fact: Duplicates are like that one song that gets stuck in your head - they keep showing up everywhere, even when you think you’ve gotten rid of them all.
Reference:
df.duplicated()- Check for duplicate rowsdf.drop_duplicates()- Remove duplicate rowsdf.drop_duplicates(subset=['col1', 'col2'])- Remove duplicates in specific columnsdf.drop_duplicates(keep='first')- Keep first occurrence of duplicates
Example:
# Check for duplicatesdf = pd.DataFrame({'A': [1, 2, 2, 3], 'B': [4, 5, 5, 6]})print(df.duplicated().sum()) # Number of duplicate rows
# Remove duplicatesdf_clean = df.drop_duplicates()print(df_clean) # Removed duplicate rowsReplacing Values
Section titled “Replacing Values”The replace() method provides a flexible way to substitute specific values or patterns in your data.
Think of replace() as find-and-replace for your data - but way more powerful than Word’s version!
Reference:
df.replace(old, new)- Replace single valuedf.replace([val1, val2], new)- Replace multiple values with same replacementdf.replace([val1, val2], [new1, new2])- Replace multiple values with different replacementsdf.replace({val1: new1, val2: new2})- Dictionary mappingdf.replace(regex=True)- Use regular expressions
Example:
# Replace sentinel values with NaNdf = pd.Series([1, -999, 2, -999, -1000, 3])df_clean = df.replace([-999, -1000], np.nan)print(df_clean) # [1.0, NaN, 2.0, NaN, NaN, 3.0]
# Different replacement for each valuedf = pd.Series(['low', 'medium', 'high', 'low'])df_mapped = df.replace({'low': 1, 'medium': 2, 'high': 3})print(df_mapped) # [1, 2, 3, 1]
# Column-specific replacement in DataFramedf = pd.DataFrame({'A': [1, 2, 3], 'B': ['x', 'y', 'z']})df = df.replace({'A': {1: 100}, 'B': {'x': 'alpha'}})print(df) # A: [100, 2, 3], B: ['alpha', 'y', 'z']Applying Custom Functions
Section titled “Applying Custom Functions”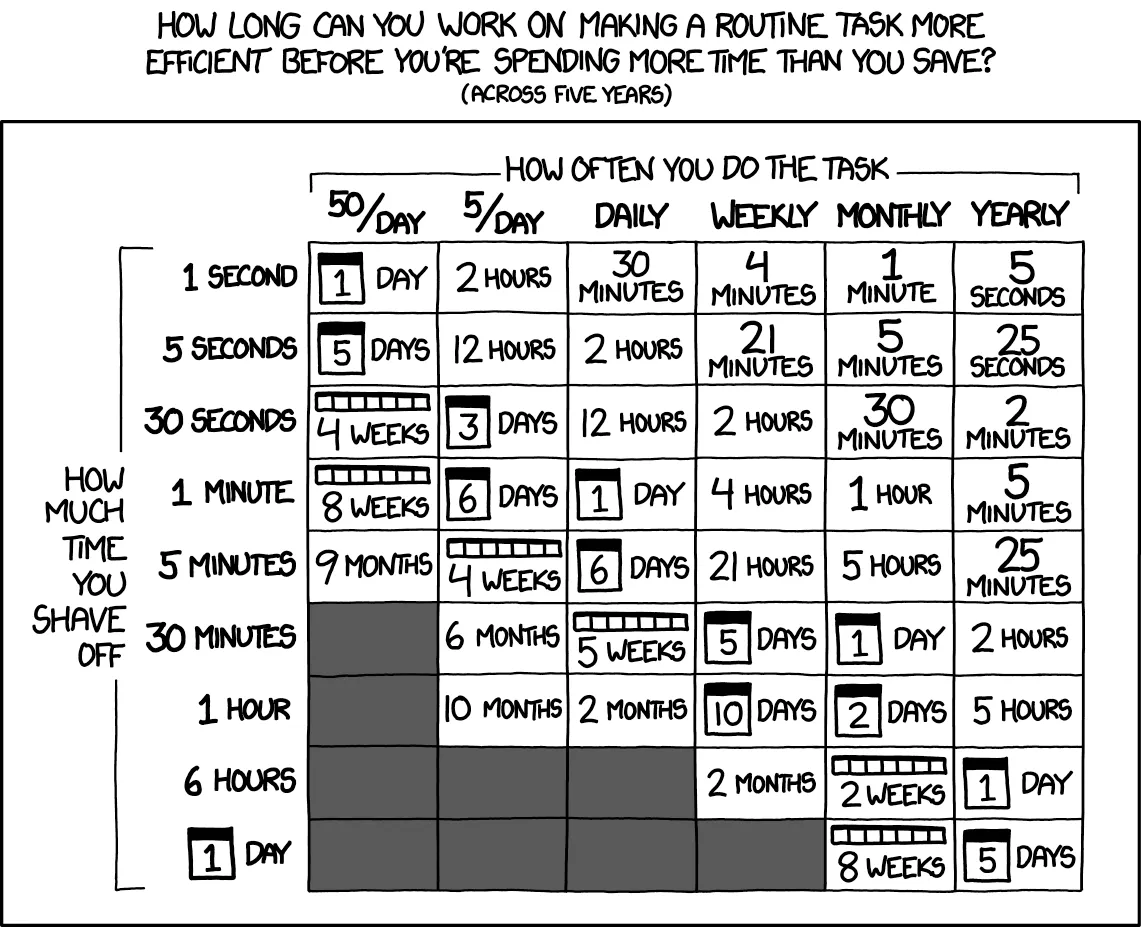 Classic time-saving calculation chart - perfect for .apply() section
Classic time-saving calculation chart - perfect for .apply() section
Sometimes built-in methods aren’t enough - you need to apply custom logic to transform your data. The .apply() and .map() methods let you use any function (built-in or custom) to transform data.
Think of .apply() as your data transformation Swiss Army knife - when pandas doesn’t have a built-in method for what you need, you can just write your own function and apply it to every row, column, or value.
Quick lambda primer: A lambda is a one-line anonymous function, perfect for simple transformations: lambda x: x * 2 is equivalent to def double(x): return x * 2, just more concise for one-time use.
Reference:
series.map(dict_or_func)- Map values in a Series (element-wise)series.apply(func)- Apply function to each element in a Seriesdf.apply(func, axis=0)- Apply function to each column (axis=0, default)df.apply(func, axis=1)- Apply function to each row (axis=1)df.map(func)- Apply function element-wise to entire DataFrame (pandas 2.1+)df.applymap(func)- Deprecated in pandas 2.1+, use.map()instead
Example:
# Clean text data with custom functiondef clean_text(text): """Remove whitespace and convert to lowercase""" return text.strip().lower()
names = pd.Series([' Alice ', 'BOB', ' Charlie'])names_clean = names.apply(clean_text)print(names_clean) # ['alice', 'bob', 'charlie']
# Map categorical values to numbersstatus = pd.Series(['active', 'inactive', 'active', 'pending'])status_map = {'active': 1, 'inactive': 0, 'pending': 2}status_coded = status.map(status_map)print(status_coded) # [1, 0, 1, 2]
# Apply function to DataFrame rowsdf = pd.DataFrame({'min': [1, 4, 7], 'max': [5, 9, 12]})df['range'] = df.apply(lambda row: row['max'] - row['min'], axis=1)print(df)# min max range# 0 1 5 4# 1 4 9 5# 2 7 12 5
# Apply function to DataFrame columnsdf = pd.DataFrame({'A': [1, 2, 3], 'B': [4, 5, 6]})column_sums = df.apply(sum, axis=0) # Sum each columnprint(column_sums) # A: 6, B: 15
# Element-wise function application (pandas 2.1+)df = pd.DataFrame({'A': [1, 2, 3], 'B': [4, 5, 6]})df_squared = df.map(lambda x: x ** 2)print(df_squared)# A B# 0 1 16# 1 4 25# 2 9 36Data Type Conversion
Section titled “Data Type Conversion”Converting data to the correct types is essential for proper analysis. This includes converting strings to numbers, dates, and other appropriate types.
Warning: Data type conversion is like trying to fit a square peg in a round hole - sometimes it works perfectly, sometimes you need to shave off a few corners, and sometimes you just need to find a different hole entirely.
Reference:
df.astype('int64')- Convert to integerdf.astype('float64')- Convert to floatdf.astype('string')- Convert to stringpd.to_datetime(df['date_column'])- Convert to datetimepd.to_numeric(df['column'], errors='coerce')- Convert to numeric, errors become NaN
Example:
# Convert data typesdf = pd.DataFrame({'A': ['1', '2', '3'], 'B': [4.5, 5.5, 6.5]})df['A'] = df['A'].astype('int64') # Convert string to integerdf['B'] = df['B'].astype('int64') # Convert float to integerprint(df.dtypes) # A: int64, B: int64Renaming Axis Indexes
Section titled “Renaming Axis Indexes”Renaming changes row or column labels without modifying data. This is essential for making your data more readable and standardizing column names.
Reference:
df.rename(index={old: new})- Rename rowsdf.rename(columns={old: new})- Rename columnsdf.rename(columns=str.lower)- Apply function to all columnsdf.rename(columns=str.strip)- Remove whitespace from column namesinplace=True- Modify DataFrame in place
Example:
# Rename specific columnsdf = pd.DataFrame({'OldName': [1, 2, 3], 'Another_Old': [4, 5, 6]})df_renamed = df.rename(columns={'OldName': 'new_name', 'Another_Old': 'better_name'})print(df_renamed.columns) # ['new_name', 'better_name']
# Apply function to all columnsdf.columns = ['First Column', ' Second ', 'THIRD']df_clean = df.rename(columns=str.lower) # Lowercase alldf_clean = df_clean.rename(columns=str.strip) # Remove spacesprint(df_clean.columns) # ['first column', 'second', 'third']
# Rename indexdf.index = ['a', 'b', 'c']df_reindexed = df.rename(index={'a': 'row_1', 'b': 'row_2'})Creating Categories
Section titled “Creating Categories”Converting continuous variables into categories makes data easier to analyze and visualize. This is especially useful for age groups, income brackets, and other meaningful categories.
Pro tip: Categories are like putting your data in organized boxes - everything has its place, and you can find things much faster when you know exactly which box to look in.
Reference:
pd.cut(series, bins)- Cut into equal-width binspd.qcut(series, q)- Cut into equal-frequency binsbins=[0, 18, 35, 50, 100]- Custom bin edgeslabels=['Young', 'Middle', 'Senior']- Custom labels for bins
Example:
# Create age groupsages = pd.Series([25, 30, 45, 60, 75])age_groups = pd.cut(ages, bins=[0, 30, 50, 100], labels=['Young', 'Middle', 'Senior'])print(age_groups) # [Young, Young, Middle, Senior, Senior]Detecting and Filtering Outliers
Section titled “Detecting and Filtering Outliers”Outliers are extreme values that may represent errors or important anomalies. Detecting and handling them appropriately is crucial for reliable analysis.

Reference:
df[df['col'] > threshold]- Filter by thresholddf.clip(lower, upper)- Cap values at boundsdf.quantile([0.25, 0.75])- Find quartiles for IQR methoddf[(df > lower) & (df < upper)]- Filter within bounds
Example:
# Remove values beyond 3 standard deviationsdf = pd.DataFrame({'value': [1, 2, 3, 100, 4, 5]})mean, std = df['value'].mean(), df['value'].std()df_clean = df[abs(df['value'] - mean) < 3 * std]print(df_clean) # Removes 100
# Cap extreme valuesdf['value'] = df['value'].clip(lower=0, upper=10)print(df) # Values capped at 0-10 range
# IQR method for outlier detectionQ1 = df['value'].quantile(0.25)Q3 = df['value'].quantile(0.75)IQR = Q3 - Q1lower_bound = Q1 - 1.5 * IQRupper_bound = Q3 + 1.5 * IQRdf_no_outliers = df[(df['value'] >= lower_bound) & (df['value'] <= upper_bound)]Categorical Data Encoding
Section titled “Categorical Data Encoding”Working with categorical data is common in data analysis. Pandas provides two main approaches: the categorical data type for efficient storage, and dummy variables for machine learning models.
Pro tip: Categorical encoding is like translating between languages - categories can be stored efficiently as codes (integers) or expanded into binary columns for models. Choose the right translation for your task!
Categorical Data Type
Section titled “Categorical Data Type”The categorical type is incredibly powerful for memory optimization, especially when you have repeated string values.
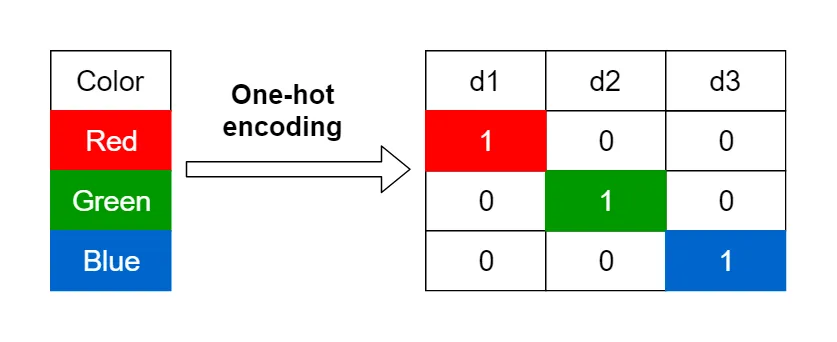 Visual showing categorical encoding: Original values → Categories → Codes with memory savings comparison
Visual showing categorical encoding: Original values → Categories → Codes with memory savings comparison
Reference:
astype('category')- Convert to categoricalcat.categories- View categoriescat.codes- View numeric codes- Use for: Repeated string values, ordered categories
Example:
# Huge memory savings for repeated valuescolors = pd.Series(['red', 'blue', 'red', 'green', 'blue'] * 1000)print(f"As object: {colors.memory_usage(deep=True)} bytes")
colors_cat = colors.astype('category')print(f"As category: {colors_cat.memory_usage(deep=True)} bytes")
# Access categories and codesprint(colors_cat.cat.categories) # ['blue', 'green', 'red']print(colors_cat.cat.codes[:5]) # [2, 0, 2, 1, 0]Creating Indicator (Dummy) Variables
Section titled “Creating Indicator (Dummy) Variables”Indicator variables convert categories into binary (0/1) columns, which is essential for machine learning models that require numeric input.
Think of dummy variables as translating categories into a language that models can understand - instead of “red”, “blue”, “green”, you get three columns of 1s and 0s indicating which color each row has.
Reference:
pd.get_dummies(series)- Create dummy variablesprefix='category'- Add prefix to column namesdrop_first=True- Avoid multicollinearity (drop first category)dtype='int64'- Specify data type for dummies (use int64 not bool - booleans can’t represent missing values)
Example:
# Create dummy variablesdf = pd.DataFrame({'color': ['red', 'blue', 'red', 'green']})dummies = pd.get_dummies(df['color'], prefix='color', dtype='int64')print(dummies)# Creates: color_blue, color_green, color_red columns with 0/1 values
# Add to original DataFramedf_with_dummies = pd.concat([df, dummies], axis=1)print(df_with_dummies)
# Drop first category to avoid multicollinearitydummies = pd.get_dummies(df['color'], prefix='color', drop_first=True, dtype='int64')print(dummies) # Only color_green and color_red (blue is the reference)LIVE DEMO! (Demo 2: Transformations - categorical encoding, string operations, sampling)
String Manipulation
Section titled “String Manipulation”Pro tip: The .str accessor is like having a Swiss Army knife for text data. It can split, join, replace, extract, and transform text in ways that would make a regex wizard jealous.
Basic String Operations
Section titled “Basic String Operations”String operations are essential for cleaning text data. Pandas provides easy-to-use string methods that work on Series containing text.
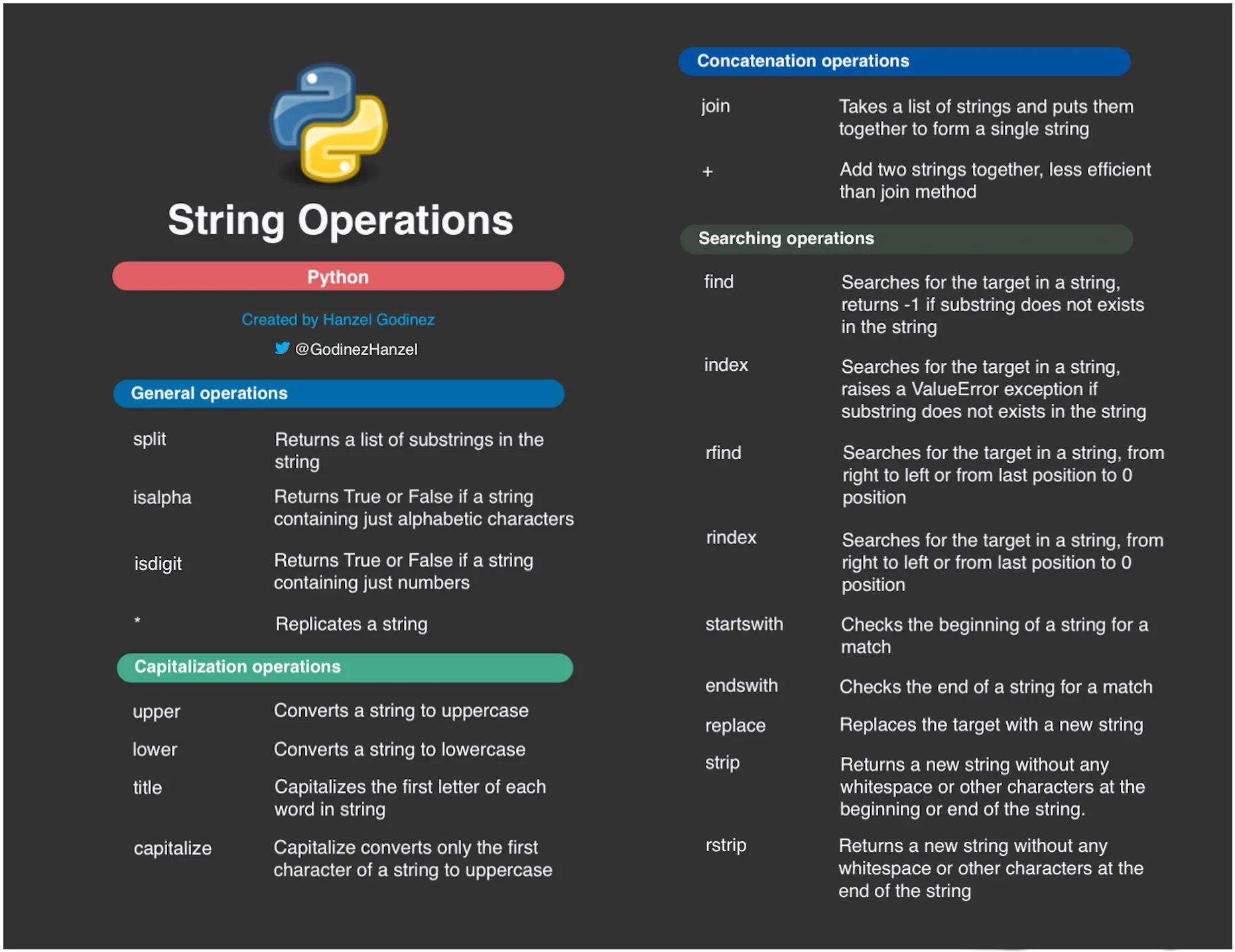 Quick reference card for common string operations: .upper()/.lower(), .strip()/.replace(), .split()/.contains()
Quick reference card for common string operations: .upper()/.lower(), .strip()/.replace(), .split()/.contains()
Input: " Alice Smith " │ ├─ .strip() ────────────► "Alice Smith" │ │ │ ├─ .lower() ────────► "alice smith" │ │ │ │ │ └─ .replace(' ', '_') ──► "alice_smith" │ │ │ └─ .split(' ') ────────► ['Alice', 'Smith'] │ │ │ └─ [0] ──────────► 'Alice' │ └─ .title() ────────────► "Alice Smith" “I got 99 problems, so I used regex. Now I have 100 problems.” - Perfect humor for string manipulation complexity
“I got 99 problems, so I used regex. Now I have 100 problems.” - Perfect humor for string manipulation complexity
Reference:
series.str.upper()- Convert to uppercaseseries.str.lower()- Convert to lowercaseseries.str.strip()- Remove leading/trailing whitespaceseries.str.replace(old, new)- Replace substringsseries.str.contains(pattern)- Check if string contains patternseries.str.startswith(prefix)- Check if string starts with prefixseries.str.endswith(suffix)- Check if string ends with suffix
Example:
# Clean text datanames = pd.Series([' Alice ', 'bob', 'CHARLIE'])clean_names = names.str.strip().str.title()print(clean_names) # ['Alice', 'Bob', 'Charlie']
# Check patternsemails = pd.Series(['alice@example.com', 'bob@test.org'])has_gmail = emails.str.contains('gmail')print(has_gmail) # [False, False]String Splitting and Joining
Section titled “String Splitting and Joining”Splitting and joining strings is common when working with structured text data like addresses, names, or delimited values.
Reference:
series.str.split(sep)- Split strings by separatorseries.str.split(sep, expand=True)- Split into separate columnsseries.str.cat(sep=' ')- Join strings with separatorseries.str.join(sep)- Join list elements with separator
Example:
# Split stringsfull_names = pd.Series(['Alice Smith', 'Bob Jones', 'Charlie Brown'])names_split = full_names.str.split(' ')print(names_split) # [['Alice', 'Smith'], ['Bob', 'Jones'], ['Charlie', 'Brown']]
# Split into columnsnames_df = full_names.str.split(' ', expand=True)print(names_df) # Two columns with first and last namesRandom Sampling and Permutation
Section titled “Random Sampling and Permutation”Random Sampling
Section titled “Random Sampling”Random sampling creates representative subsets of data for analysis, testing, and machine learning. It’s essential for creating train/test splits, bootstrap analysis, and data exploration.
Reference:
df.sample(n=None, frac=None, replace=False, weights=None, random_state=None)- Random samplingn=10- Sample exactly 10 rowsfrac=0.5- Sample 50% of rowsreplace=True- Sample with replacement (bootstrap)weights='column'- Weighted sampling by column valuesrandom_state=42- Reproducible samplingdf.iloc[::step]- Systematic sampling every nth row
Example:
# Random samplingdf = pd.DataFrame({'A': range(100), 'B': range(100, 200)})sample = df.sample(n=10, random_state=42) # Sample 10 rowsprint(len(sample)) # 10
# Stratified samplingdf['category'] = ['A', 'B'] * 50stratified = df.groupby('category').apply(lambda x: x.sample(2))print(len(stratified)) # 4 (2 from each category)Permutation and Shuffling
Section titled “Permutation and Shuffling”Permutation randomizes data order while preserving relationships. It’s essential for cross-validation, bootstrap analysis, and breaking temporal dependencies in time series data.
Reference:
df.sample(frac=1)- Shuffle all rows (permutation)df.reindex(np.random.permutation(df.index))- Permute index orderdf.sample(n=len(df), replace=True)- Bootstrap samplingnp.random.permutation(array)- Randomly permute arrayrandom_state=42- Reproducible permutation
Example:
# Shuffle DataFramedf = pd.DataFrame({'A': [1, 2, 3, 4], 'B': [5, 6, 7, 8]})shuffled = df.sample(frac=1, random_state=42)print(shuffled) # Random order of rows
# Bootstrap samplingbootstrap = df.sample(n=len(df), replace=True, random_state=42)print(len(bootstrap)) # 4 (same length, but with replacement)Data Validation and Quality Assessment
Section titled “Data Validation and Quality Assessment” Shows data errors invalidating research - perfect for validation section
Shows data errors invalidating research - perfect for validation section
| Issue | Detection | Solution |
|---|---|---|
| Missing Values | df.isnull().sum() | • Impute (mean/median) • Forward/backward fill • Drop if <5% missing |
| Duplicates | df.duplicated() | • drop_duplicates()• Keep first or last |
| Wrong Data Type | df.dtypes | • astype('int64')• pd.to_datetime() |
| Outliers | df.describe()Box plots | • IQR method filter • Clip extreme values • Keep if valid (verify) |
| Inconsistent Categories | df['col'].unique() | • str.lower().strip()• replace() mapping |
Data Quality Checks
Section titled “Data Quality Checks”Data quality checks identify issues like missing values, duplicates, outliers, and data type inconsistencies. These checks are essential for ensuring reliable analysis results.
Reference:
df.isnull().sum()- Count missing values per columndf.duplicated().sum()- Count duplicate rowsdf.nunique()- Count unique values per columndf.dtypes- Data types per columndf.describe()- Summary statistics (numeric columns only by default)df.describe(include='all')- Summary statistics for all columns (numeric + categorical)df.describe(include=['object'])- Summary statistics for categorical columns onlydf.info()- Detailed informationdf.memory_usage()- Memory usage per column
Example:
# Data quality assessmentdf = pd.DataFrame({'A': [1, 2, 2, 4], 'B': [5, 6, 6, 8], 'C': [9, 10, 11, 12]})print(df.isnull().sum()) # Missing values per columnprint(df.duplicated().sum()) # Number of duplicate rowsprint(df.nunique()) # Unique values per columnprint(df.dtypes) # Data types per columnData Validation Rules
Section titled “Data Validation Rules”Data validation rules ensure data meets business requirements and constraints. These rules help maintain data integrity and prevent analysis errors.
Reference:
df[condition]- Filter rows meeting conditiondf.between(left, right)- Check if values are between boundsdf.isin(values)- Check if values are in listdf.str.contains(pattern)- Check if strings contain patterndf.str.match(pattern)- Check if strings match patterndf.str.len()- Get string lengthdf.str.isdigit()- Check if strings are digits
Example:
# Data validation rulesdf = pd.DataFrame({'Age': [25, 30, 35, 40], 'Email': ['alice@test.com', 'bob@example.org', 'charlie@test.com', 'diana@example.org']})
# Age validation (18-65)valid_ages = df[df['Age'].between(18, 65)]print(valid_ages) # All rows (ages are valid)
# Email validationemail_pattern = r'^[a-zA-Z0-9._%+-]+@[a-zA-Z0-9.-]+\.[a-zA-Z]{2,}$'valid_emails = df[df['Email'].str.match(email_pattern)]print(valid_emails) # All rows (emails are valid)Data Cleaning Pipeline
Section titled “Data Cleaning Pipeline”A systematic approach to data cleaning ensures consistent, high-quality results. Follow these steps in order for best results.
graph TD A[Load Data] --> B{Inspect Data} B --> C[Check Missing Values] B --> D[Check Duplicates] B --> E[Check Data Types] B --> F[Check Outliers]
C --> G{Issues Found?} D --> G E --> G F --> G
G -->|Yes| H[Handle Issues] G -->|No| L[Validate Results]
H --> I[Fill/Drop Missing] H --> J[Remove Duplicates] H --> K[Convert Types] H --> M[Handle Outliers]
I --> L J --> L K --> L M --> L
L --> N{Data Quality OK?} N -->|No| B N -->|Yes| O[Document Decisions] O --> P[Export Clean Data]Think of data cleaning as being a detective - you need to follow the clues, ask the right questions, and sometimes you have to make tough decisions about what to keep and what to throw away.
Reference:
- Load and inspect data -
df.head(),df.info(),df.describe() - Handle missing values -
df.isnull().sum(),df.fillna(),df.dropna() - Remove duplicates -
df.duplicated(),df.drop_duplicates() - Convert data types -
df.astype(),pd.to_datetime(),pd.to_numeric() - Handle outliers -
df.quantile(),df.clip(), filtering - Validate data quality - Check ranges, patterns, consistency
- Export clean data -
df.to_csv(),df.to_excel()
Example:
# Step 1: Load and inspectdf = pd.read_csv('messy_data.csv')print(df.info())print(df.isnull().sum())
# Step 2: Handle missing valuesdf['Age'].fillna(df['Age'].mean(), inplace=True)df['Name'].fillna('Unknown', inplace=True)
# Step 3: Remove duplicatesdf = df.drop_duplicates()
# Step 4: Convert data typesdf['Age'] = df['Age'].astype('int64')df['Date'] = pd.to_datetime(df['Date'])
# Step 5: Handle outliersQ1, Q3 = df['Salary'].quantile([0.25, 0.75])IQR = Q3 - Q1df = df[~((df['Salary'] < Q1 - 1.5*IQR) | (df['Salary'] > Q3 + 1.5*IQR))]
# Step 6: Validateprint(df.describe())print(df.dtypes)
# Step 7: Exportdf.to_csv('clean_data.csv', index=False)Configuration-Driven Processing
Section titled “Configuration-Driven Processing”Configuration files make data cleaning pipelines more maintainable and reproducible. Simple text files or Python dictionaries can store filter rules, cleaning parameters, and processing steps.
Pro tip: Keep your data cleaning logic separate from your parameters. This makes your code more maintainable and your pipelines more reproducible.
Reference:
- Use Python dictionaries for simple configurations
- Store parameters in separate files (CSV, JSON, or simple text)
- Keep cleaning logic in functions
- Document your cleaning decisions
Example:
# Simple configuration dictionarycleaning_config = { 'missing_strategies': { 'age': 'median', 'income': 'mean', 'date': 'ffill' }, 'outlier_threshold': 3.0, 'drop_columns': ['temp_id', 'notes']}
# Apply configurationfor column, strategy in cleaning_config['missing_strategies'].items(): if strategy == 'median': df[column].fillna(df[column].median(), inplace=True) elif strategy == 'mean': df[column].fillna(df[column].mean(), inplace=True) elif strategy == 'ffill': df[column].fillna(method='ffill', inplace=True)Running Notebooks from Command Line
Section titled “Running Notebooks from Command Line”For automated pipelines and batch processing, you can execute Jupyter notebooks from the command line without opening the Jupyter interface.
Basic Execution
Section titled “Basic Execution”# Execute a single notebookjupyter nbconvert --execute --to notebook your_notebook.ipynb
# Execute and save output to a new filejupyter nbconvert --execute --to notebook --output executed_notebook your_notebook.ipynb
# Execute and overwrite the original filejupyter nbconvert --execute --to notebook --inplace your_notebook.ipynbNotebook Pipeline Automation
Section titled “Notebook Pipeline Automation”Always check “exit codes” after notebook execution to ensure your pipeline stops if any step fails. When a command runs successfully it returns an exit code of 0, other values (usually 1) indicate an error.
You may check exit codes using the special variable $?, which contains exit code for the previous command. Alternatively, we can use an OR operator (||) to instruct the shell to do something when a command fails.
Note: The || operator means “OR” - if the command fails (non-zero exit code), execute the code block in curly braces {}. This is more concise than checking $? explicitly.
#!/bin/bash# Example pipeline script
echo "Starting data analysis pipeline..."
# Run notebooks in sequencejupyter nbconvert --execute --to notebook q4_exploration.ipynbif [ $? -ne 0 ]; then echo "ERROR: Q4 exploration failed" exit 1fi
jupyter nbconvert --execute --to notebook q5_missing_data.ipynb || { echo "ERROR: Q5 missing data analysis failed" exit 1}
echo "Pipeline completed successfully!"Key Parameters
Section titled “Key Parameters”--execute: Run all cells in the notebook--to notebook: Keep output as notebook format--inplace: Overwrite the original file--output filename: Save to a new file--allow-errors: Continue execution even if cells fail
LIVE DEMO!
Section titled “LIVE DEMO!”(Demo 3: Complete Workflow - end-to-end data cleaning pipeline)