08: Data Wrangling
See Bonus for advanced topics:
- Advanced groupby operations with custom functions
- Hierarchical grouping and MultiIndex operations
- Performance optimization for large datasets
- Custom aggregation functions and transformations
- Advanced pivot table operations
Outline
Section titled “Outline”- groupby split-apply-combine essentials
- pivot tables and crosstab basics
- remote workflows: ssh, screen, tmux
- performance-minded patterns beginners should know
Fun fact: The term “aggregation” comes from the Latin “aggregare” meaning “to add to a flock.” In data science, we’re literally gathering scattered data points into meaningful groups - turning a flock of individual observations into organized insights.
Data aggregation is the process of summarizing and grouping data to extract meaningful insights. This lecture covers the essential tools for data aggregation: groupby operations, pivot tables, and remote computing for handling large datasets.
The Split-Apply-Combine Paradigm
Section titled “The Split-Apply-Combine Paradigm”Reality check: GroupBy operations are the bread and butter of data analysis. Master this concept and you’ll be able to answer almost any “what if we group by…” question that comes your way.
The split-apply-combine paradigm is the foundation of data aggregation. You split data into groups, apply a function to each group, and combine the results.
Visual Guide - GroupBy Operations:
BEFORE GROUPBY AFTER GROUPBY┌─────────┬─────────┬─────────┐ ┌─────────┬─────────┐│ Category│ Value │ Other │ │ Category│ Mean │├─────────┼─────────┼─────────┤ ├─────────┼─────────┤│ A │ 10 │ X │ │ A │ 10.0 ││ A │ 15 │ Y │ │ B │ 25.0 ││ B │ 20 │ Z │ └─────────┴─────────┘│ B │ 25 │ W ││ A │ 5 │ V ││ B │ 30 │ U │└─────────┴─────────┴─────────┘Visual Guide - Split-Apply-Combine:
ORIGINAL DATA SPLIT BY CATEGORY┌─────────┬─────────┬─────────┐ ┌─────────┬─────────┐│ Category│ Value │ Other │ │ Group A │ Group B │├─────────┼─────────┼─────────┤ ├─────────┼─────────┤│ A │ 10 │ X │ │ A, 10 │ B, 20 ││ A │ 15 │ Y │ │ A, 15 │ B, 25 ││ B │ 20 │ Z │ │ A, 5 │ B, 30 ││ B │ 25 │ W │ └─────────┴─────────┘│ A │ 5 │ V ││ B │ 30 │ U │└─────────┴─────────┴─────────┘
APPLY FUNCTION (e.g., mean) COMBINE RESULTS┌─────────┬─────────┐ ┌─────────┬─────────┐│ Group A │ Group B │ │ Category│ Mean │├─────────┼─────────┤ ├─────────┼─────────┤│ mean(10,│ mean(20,│ │ A │ 10.0 ││ 15, 5) │ 25, 30)│ │ B │ 25.0 ││ = 10.0 │ = 25.0 │ └─────────┴─────────┘└─────────┴─────────┘Basic GroupBy Operations
Section titled “Basic GroupBy Operations”Reference:
df.groupby('column')- Group by single columndf.groupby(['col1', 'col2'])- Group by multiple columnsgrouped.mean()- Calculate mean for each groupgrouped.sum()- Calculate sum for each groupgrouped.count()- Count non-null valuesgrouped.size()- Count all values (including nulls)grouped.agg(['mean', 'sum', 'count'])- Multiple aggregations
Example:
import pandas as pdimport numpy as np
# Create sample datadf = pd.DataFrame({ 'Department': ['Sales', 'Sales', 'Engineering', 'Engineering'], 'Employee': ['Alice', 'Bob', 'Charlie', 'Diana'], 'Salary': [50000, 55000, 80000, 85000], 'Experience': [2, 3, 5, 7]})
# Basic groupby operationsprint("Group by Department:")print(df.groupby('Department')['Salary'].mean())
print("\nMultiple aggregations:")print(df.groupby('Department').agg({ 'Salary': ['mean', 'sum'], 'Experience': 'mean'}))Advanced GroupBy Operations
Section titled “Advanced GroupBy Operations”Transform Operations
Section titled “Transform Operations”Transform operations apply a function to each group and return a result with the same shape as the original data.
Reference:
grouped.transform('mean')- Apply mean to each groupgrouped.transform('std')- Apply standard deviation to each groupgrouped.transform(lambda x: x - x.mean())- Custom transform functiongrouped.transform(['mean', 'std'])- Multiple transforms
Example:
# Transform: Add group means as new columndf['Salary_Mean'] = df.groupby('Department')['Salary'].transform('mean')df['Salary_Std'] = df.groupby('Department')['Salary'].transform('std')df['Salary_Normalized'] = df.groupby('Department')['Salary'].transform(lambda x: (x - x.mean()) / x.std())
print("Data with group statistics:")print(df[['Department', 'Employee', 'Salary', 'Salary_Mean', 'Salary_Std', 'Salary_Normalized']])Filter Operations
Section titled “Filter Operations”Filter operations remove entire groups based on a condition.
Reference:
grouped.filter(lambda x: len(x) > n)- Keep groups with more than n rowsgrouped.filter(lambda x: x['col'].sum() > threshold)- Keep groups meeting conditiongrouped.filter(lambda x: x['col'].mean() > threshold)- Filter by group statistics
Example:
# Filter: Keep only departments with more than 1 employeefiltered = df.groupby('Department').filter(lambda x: len(x) > 1)print("Departments with multiple employees:")print(filtered)
# Filter: Keep only departments with average salary > 60000high_salary_depts = df.groupby('Department').filter(lambda x: x['Salary'].mean() > 60000)print("\nHigh-salary departments:")print(high_salary_depts)Apply Operations
Section titled “Apply Operations”Apply operations let you use custom functions on each group.
Reference:
grouped.apply(func)- Apply custom function to each groupgrouped.apply(lambda x: x.sort_values('col'))- Sort each groupgrouped.apply(lambda x: x.nlargest(2, 'col'))- Get top 2 from each groupgrouped.apply(func, include_groups=False)- Exclude grouping columns from function (pandas 2.2+)
Important: FutureWarning for include_groups Parameter
Starting in pandas 2.2, when using .apply() on a GroupBy object, pandas will include the grouping columns in the DataFrame passed to your function. This is a change from previous behavior where grouping columns were excluded. To maintain the old behavior (where grouping columns are excluded), you should explicitly set include_groups=False.
What’s happening:
- Old behavior (pandas < 2.2): When you call
df.groupby('Department').apply(func), the function receives only the non-grouping columns - New behavior (pandas 2.2+): By default, the function receives all columns including the grouping columns
- Future behavior:
include_groups=Falsewill become the default, but you should explicitly set it now to avoid warnings
Why this matters:
- If your function expects only non-grouping columns, you’ll get unexpected behavior
- The warning helps you prepare for future pandas versions
- Setting
include_groups=Falseexplicitly makes your code future-proof
Example:
# Apply: Custom function for salary statisticsdef salary_stats(group): # With include_groups=False, 'group' contains only non-grouping columns # Without it, 'group' also contains 'Department' column return pd.Series({ 'count': len(group), 'mean': group['Salary'].mean(), 'std': group['Salary'].std(), 'range': group['Salary'].max() - group['Salary'].min() })
print("Custom statistics by department:")# Explicitly set include_groups=False to avoid FutureWarningprint(df.groupby('Department').apply(salary_stats, include_groups=False))
# Apply: Get top earners in each departmenttop_earners = df.groupby('Department').apply( lambda x: x.nlargest(1, 'Salary'), include_groups=False)print("\nTop earners per department:")print(top_earners)LIVE DEMO!
Section titled “LIVE DEMO!”Hierarchical Grouping
Section titled “Hierarchical Grouping”Reference:
df.groupby(['level1', 'level2'])- Multi-level groupinggrouped.unstack()- Convert to wide formatgrouped.stack()- Convert to long formatgrouped.swaplevel(0, 1)- Swap grouping levels
Example:
# Create hierarchical datahierarchical_df = pd.DataFrame({ 'Region': ['North', 'North', 'South', 'South', 'North', 'South'], 'Department': ['Sales', 'Engineering', 'Sales', 'Engineering', 'Marketing', 'Marketing'], 'Revenue': [100000, 150000, 120000, 180000, 80000, 90000], 'Employees': [5, 8, 6, 10, 4, 5]})
# Hierarchical groupinghierarchical_grouped = hierarchical_df.groupby(['Region', 'Department']).sum()print("Hierarchical grouping:")print(hierarchical_grouped)
# Unstack to wide formatwide_format = hierarchical_grouped.unstack()print("\nWide format:")print(wide_format)Pivot Tables and Cross-Tabulations
Section titled “Pivot Tables and Cross-Tabulations”
Think of pivot tables as the data analyst’s Swiss Army knife - they can reshape, summarize, and analyze data in ways that would take dozens of lines of code to accomplish manually.
Pivot tables are powerful tools for summarizing and analyzing data across multiple dimensions.
Visual Guide - Pivot Table Transformation:
LONG FORMAT (Original) WIDE FORMAT (Pivoted)┌─────────┬─────────┬─────────┐ ┌─────────┬─────────┬─────────┐│ Product │ Region │ Sales │ │ Product │ North │ South │├─────────┼─────────┼─────────┤ ├─────────┼─────────┼─────────┤│ A │ North │ 1000 │ │ A │ 1000 │ 1500 ││ A │ South │ 1500 │ │ B │ 2000 │ 1200 ││ B │ North │ 2000 │ └─────────┴─────────┴─────────┘│ B │ South │ 1200 │└─────────┴─────────┴─────────┘Basic Pivot Tables
Section titled “Basic Pivot Tables”Reference:
pd.pivot_table(df, values='col', index='row', columns='col')- Basic pivotpd.pivot_table(df, aggfunc='mean')- Specify aggregation functionpd.pivot_table(df, fill_value=0)- Fill missing valuespd.pivot_table(df, margins=True)- Add totalspd.crosstab(index, columns)- Cross-tabulation
Example:
# Create sample sales datasales_data = pd.DataFrame({ 'Product': ['A', 'A', 'B', 'B', 'C', 'C'], 'Region': ['North', 'South', 'North', 'South', 'North', 'South'], 'Sales': [1000, 1500, 2000, 1200, 800, 900]})
# Basic pivot tablepivot = pd.pivot_table(sales_data, values='Sales', index='Product', columns='Region', aggfunc='sum')print("Sales by Product and Region:")print(pivot)
# Pivot with multiple aggregationspivot_multi = pd.pivot_table(sales_data, values='Sales', index='Product', columns='Region', aggfunc=['sum', 'mean'])print("\nMultiple aggregations:")print(pivot_multi)Advanced Pivot Operations
Section titled “Advanced Pivot Operations”Reference:
pivot_table(..., margins=True, margins_name='Total')- Add totalspivot_table(..., fill_value=0)- Fill missing valuespivot_table(..., dropna=False)- Keep missing combinationspivot_table(..., observed=True)- Include all category combinations
Example:
# Advanced pivot with totals and missing value handlingadvanced_pivot = pd.pivot_table(sales_data, values='Sales', index='Product', columns='Region', aggfunc='sum', margins=True, margins_name='Total', fill_value=0)print("Advanced pivot with totals:")print(advanced_pivot)
# Cross-tabulationcrosstab = pd.crosstab(sales_data['Product'], sales_data['Region'], margins=True)print("\nCross-tabulation:")print(crosstab)LIVE DEMO!
Section titled “LIVE DEMO!”Remote Computing and SSH
Section titled “Remote Computing and SSH”
When your data is too big for your laptop, it’s time to think about remote computing. SSH is your gateway to powerful remote servers that can handle massive datasets.
Remote computing allows you to leverage powerful servers for data analysis that would be impossible on your local machine.
SSH Fundamentals
Section titled “SSH Fundamentals”Reference:
ssh username@hostname- Connect to remote serverssh -p port username@hostname- Connect on specific portssh-keygen -t rsa- Generate SSH key pairssh-copy-id username@hostname- Copy public key to serverscp file username@hostname:path- Copy file to serverscp username@hostname:file path- Copy file from server
Example:
# Generate SSH key pairssh-keygen -t rsa -b 4096 -C "your_email@example.com"
# Copy public key to serverssh-copy-id username@server.com
# Connect to serverssh username@server.com
# Copy files to serverscp data.csv username@server.com:~/data/
# Copy files from serverscp username@server.com:~/results/analysis.ipynb ./Remote Data Analysis
Section titled “Remote Data Analysis”Reference:
# Start Jupyter notebook on remote serverjupyter notebook --ip=0.0.0.0 --port=8888 --no-browser
# Forward port to local machinessh -L 8888:localhost:8888 username@server.com
# Access Jupyter at http://localhost:8888# As if it were running on your local machineExample:
# Remote data analysis workflowimport pandas as pdimport numpy as np
# Load large dataset on remote serverdf = pd.read_csv('/path/to/large_dataset.csv')
# Perform aggregation on remote serverresult = df.groupby('category').agg({ 'value': ['mean', 'std', 'count'], 'other_col': 'sum'})
# Save resultsresult.to_csv('aggregated_results.csv')
# Download results to local machine# scp username@server.com:~/aggregated_results.csv ./screen and tmux for Persistent Sessions
Section titled “screen and tmux for Persistent Sessions”
Screen lets you detach and reattach long-running jobs; tmux is a more modern, scriptable alternative. Use whichever your server offers.
Screen quickstart:
# Create a named screen sessionscreen -S analysis
# Detach (Ctrl+a d) and list sessionsscreen -ls
# Reattach laterscreen -r analysis
# Kill session from insideexittmux quickstart:
Reference:
# tmux commandstmux new-session -s analysistmux list-sessionstmux attach-session -t analysistmux kill-session -t analysis
# Inside tmuxCtrl+b d # Detach from sessionCtrl+b c # Create new windowCtrl+b n # Next windowCtrl+b p # Previous windowExample:
# Start persistent analysis sessiontmux new-session -s data_analysis
# Inside tmux, start your analysisconda activate datasci_217jupyter notebook --ip=0.0.0.0 --port=8888
# Detach from session (Ctrl+b, then d)# Session continues running on server
# Reattach latertmux attach-session -t data_analysisPerformance Optimization
Section titled “Performance Optimization”
When working with large datasets, every millisecond counts. Understanding performance optimization can mean the difference between a 5-minute analysis and a 5-hour analysis.
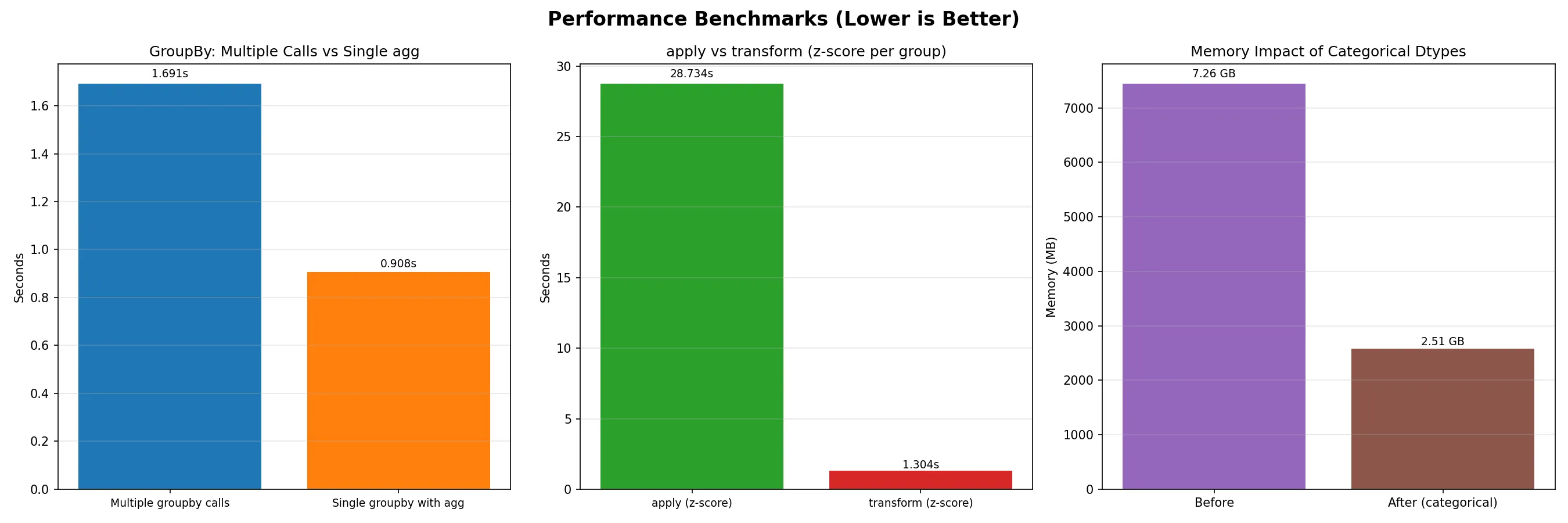
Efficient GroupBy Operations
Section titled “Efficient GroupBy Operations”Reference:
# Optimize groupby operationsdef efficient_groupby(df, group_cols, agg_cols): """Efficient groupby with optimized operations"""
# Use categorical data types for grouping columns for col in group_cols: if df[col].dtype == 'object': df[col] = df[col].astype('category')
# Use specific aggregation functions result = df.groupby(group_cols)[agg_cols].agg({ 'numeric_col': ['mean', 'sum'], 'other_col': 'count' })
return result
# Memory-efficient operations## Note: chunking manually for larger-than-memory data should be a last resort.## It is usually faster to rely on package-provided options.def memory_efficient_analysis(df): """Analyze large dataset with chunking"""
# Process in chunks chunk_size = 10000 results = []
for chunk in pd.read_csv('large_file.csv', chunksize=chunk_size): # Process chunk chunk_result = chunk.groupby('category').sum() results.append(chunk_result)
# Combine results final_result = pd.concat(results).groupby(level=0).sum() return final_resultParallel Processing (optional)
Section titled “Parallel Processing (optional)”Reference:
from multiprocessing import Poolimport pandas as pd
def process_chunk(chunk): """Process a single chunk of data""" return chunk.groupby('category').sum()
def parallel_groupby(df, n_processes=4): """Parallel groupby processing"""
# Split data into chunks chunk_size = len(df) // n_processes chunks = [df.iloc[i:i+chunk_size] for i in range(0, len(df), chunk_size)]
# Process in parallel with Pool(n_processes) as pool: results = pool.map(process_chunk, chunks)
# Combine results return pd.concat(results).groupby(level=0).sum()